"Silent mutations" can change cancer cells
Gene mutations in the protein blueprint were previously considered irrelevant if the same protein building blocks are still produced. Now researchers from Freiburg have shown that these mutations can indeed alter protein activity in cancer cells.
Until now, it was thought that changes in the genome had no consequences if they did not lead to an exchange of protein building blocks. This was referred to as "silent mutations". Now researchers from Freiburg have shown that such changes can indeed alter cell activity. The team led by Prof. Dr. Sven Diederichs, who heads the Department of Oncological Research at the Department of Thoracic Surgery at the Freiburg University Medical Center and the Department of RNA Biology and Cancer at the German Cancer Research Center (DKFZ), investigated 88 tumor types and more than 650,000 mutations. "Using the important cancer gene KRAS as an example, we were able to show that a supposedly silent mutation reduces protein formation. So silent mutations are not so silent after all," says Diederichs. The researchers developed a user-friendly database, SynMICdb, which lists known silent mutations and makes it much easier for other scientists to analyze them. The researchers will present their database and concrete results on June 12, 2019 in the journal Nature Communications .
Like a misfolded city map
G, C, A, T: there are four bases in the genome, each arranged in groups of three. This allows 64 units of information to be represented. However, the proteins in the human body are made up of only 21 different amino acids. Therefore, for many proteins there are several synonymous ways of representing them in the genome. These changes have long been referred to as "silent mutations". Using the example of the oncogene KRAS, the researchers showed that even a single synonymous mutation changes the structure of the genetic material transcript mRNA. This can make the mRNA more difficult to read and the protein is produced less.
"It's a bit like a city map: Depending on how it is folded, you can read it better or worse. How quickly you get to your destination depends on this. The map is the same, but the folding has consequences," explains Diederichs.
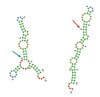
Bioinformaticians at the University of Freiburg were also able to simulate such a misfolding of the mRNA on the computer. "We were able to show that the spatial properties can change dramatically even in the case of mutations that are actually silent," says Prof. Dr. Rolf Backofen from the Institute of Computer Science at the University of Freiburg.
"Such changes could play an important role in cancer therapy in the future, for example because crucial proteins are produced more or less strongly. Before this can happen, however, the consequences of synonymous mutations need to be much better researched. Thanks to our online database, this is now also possible for scientists without in-depth bioinformatics knowledge," says Diederichs.
Picture caption: Wrongly folded: A single silent mutation causes the structure of the protein blueprint to change.
Image source: Freiburg University Medical Center
Original title of the study: A pan-cancer analysis of synonymous mutations
DOI: 10.1038/s41467-019-10489-2
Link to the study: https: //www.nature.com/articles/s41467-019-10489-2
Link to the database: synmicdb.dkfz.de
Contact:
Prof. Dr. Sven Diederichs
Head of the Department of Oncological Research
Department of Thoracic Surgery
Freiburg University Medical Center
and
Head of the Department of RNA Biology and Cancer
German Cancer Research Center (DKFZ) Heidelberg
Phone: 0761 270-77571
sven.diederichs@uniklinik-freiburg.de
Back
Medical Center - University of Freiburg
Central Information
Phone: 0761 270-0
info@uniklinik-freiburg.de
For press inquiries:
Corporate Communications
Breisacher Straße 153
79110 Freiburg
Phone: 0761 270-84830
kommunikation@uniklinik-freiburg.de