Digital PCR
dPCR
Digital PCR is a form of quantitative PCR based on end-point measurements. It has distinct advantages over QPCR for specific types of assays:
- Absolute quantification
- Detection of rare target sequences
- Single cell analysis
- Detection of small fold changes (+/-10%)
Lighthouse has a recently acquired Stilla Naica Crystal Digital PCR System available for you to use. Specific differences between the two systems can be found below.
What is Digital PCR?
Digital PCR excels in the measurement of low copy numbers, and has a linear range of 5 logs (from 1 to approximately 100,000 copies per reaction). The linearity of the process is what allows the detection of small-fold changes.
Because of its ability to absolutely quantify the number of molecules present within a sample, the use of a reference gene is not an absolute requirement in digital PCR. In addition, dPCR allows rare molecules to be detected within complex mixtures much more efficiently than standard QPCR. This is particularly useful in such applications as detecting circulating tumor DNA in cancer patients, where there are overall very few molecules of DNA to detect, within a large background of wild type DNA.
Digital PCR: Lighthouse System Comparisons
| System and Detection Method | Fluorophores | Hydrolysis (Taqman) Probes | Intercalating Dyes (EvaGreen) | Additional Reaction Components Needed | Examples of Compatible Mixes |
Stilla Naica (Probes) | FAM or FAM | yes | - | Fluorescein
| Probes mixes: |
Stilla Naica (EvaGreen) | EvaGreen | - | yes | EvaGreen dye (20x in water) Alexa Fluor 647 Dextran | EvaGreen mix:
|
Digital PCR: How does it work?
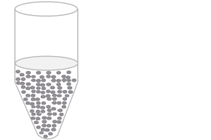
Digital PCR works by partitioning the QPCR reaction into very many small reactions (less than approx. 1 nL volume). Lighthouse systems create many small droplets within a water-in-oil emulsion, located on a special optical slide (Stilla Naica).
Droplets are generated as an emulsion in which the sample is partitioned into up to 25,000 aqueous droplets within an oil carrier.
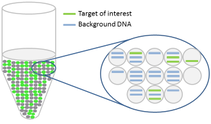
The target and background DNA are randomly distributed throughout the droplets as the droplets are formed. The sample can then be run on an ordinary thermal cycler.
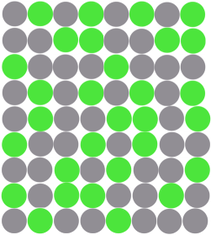
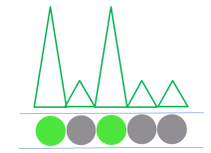
After the PCR amplification step, each droplet provides a fluorescent positive or negative signal indicating whether the target was present or not within the droplet after partitioning.
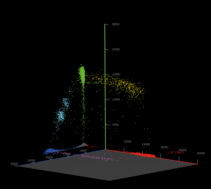
After cycling, the sample is placed on the Droplet Reader where the droplets are examined individually, providing an independent digital measurement. Depending on their signal, the droplets are scored as either positive or negative for the marker. Based on the number of positive droplets and the number of partitions/droplets overall, the number of target molecules which were originally present, in the reaction. can be calculated ith the help of the Poisson distribution.
Stilla Naica System - Digital PCR
Lighthouse has a Stilla Naica Crystal Digital PCR system.
Digital PCR (cdPCR as well as ddPCR) is related to quantitative PCR (qPCR), but has distinct advantages for certain types of assays:
- Absolute quantification
- Detection of rare target sequences
- Single cell analysis
- Detection of small fold changes (+/-10%)
The mayor differences between the Stilla system and other droplet based systems:
On the Stilla system the sample first flows through a network of microchannels and is partitioned into a large 2D array of up to 25,000 individual droplets, also called a droplet crystal. PCR is then performed on-chip and the droplets are scanned to reveal those which contain amplified targets. The last step consists of counting the number of these positive droplets to precisely extract the absolute quantity of nucleic acids.
On the Stilla system the sample first flows through a network of microchannels and is partioned into a large 2D array of up to 25,000 individual droplets, also called droplet crystal. PCR is the performed on-chip and the droplets are scanned to reveal those which contain amplified targets. The last step consists of counting the number of these droplets to prececiesely extract the absolute quantity of nucleic acids.
Workflow
- Sample pipetting: Only one pipetting step is required to load the reaction mixes into the wells of the Sapphire (or Ruby) Chips.
- Droplet crystal generation and amplification: Once the chips are placed into the Naica Geode, launch the combined partitioning and thermocycling program. Crystals of up to 25,000 droplets are created from each sample, PCR amplification is performed immediately after crystal generation.
- Crystal reading: Transfer the chips to the Naica Prism6 instrument. Crystals are read using up to 6 fluorescence channels.
- Data analysis: The crystal reader program Crystal Miner software automatically measures the concentrations of targeted nucleic acids. Analyze your data and explore your crystals using the program.
Qiagen QIAcuity Digital PCR System
Lighthouse also has a Qiagen QIAcuity available for use. The QIAcuity is a nanoplate based system which uses a grid to form partitions within the wells, rather than droplets in an emulsion. The QIAcuity is a five-color system able to look at the same assortment of fluorophores as the Stilla Naica, with the exception of Atto 700.
The Qiagen system works with 24-well or 96-well nanoplates and is compatible with standard QPCR mixes.
| Nanoplate type | Partitions per Well | Rxn. Vol. | Catalog Number |
| 96-well | 8.5k | 12 µL | 250021 |
| 24-well | 8.5k | 12 µL | 250011 |
| 24-well | 26k | 40 µL | 250001 |
When used with Actome's Protein Interaction Coupling (PICO) Kits, the QIAcuity system also enables the analysis of proteins, protein interactions, and post-translational modifications from biological systems, low background and low sample input volume. For information about this application, please contact Actome.
Further Information
Some useful documents about dPCR:
| Stilla Naica Crystal Miner User Manual v.2.0 | User manual for the Naica Crystal Miner software |
| Stilla Naica Crystal Reader User Manual 2.0 | User manual for the Naica Crystal Reader |
Bio-Rad technical note on detecting rare mutant alleles with ddPCR | |
| Bio-Rad technical note on looking at copy number variation by ddPCR |
For more information about using the machine or for help in designing possible probes/primers, please contact the Lighthouse Core Facility.
General Applications Guide
General Applications Guide for droplet digital PCR with many application descriptions and tips and tricks